Abstract
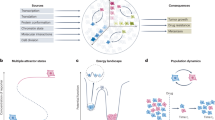
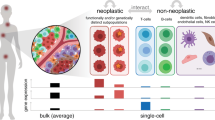
The transformation of normal cells into cancer cells and maintenance of the malignant state and phenotypes are associated with genetic and epigenetic deregulations, altered cellular signaling responses and aberrant interactions with the microenvironment. These alterations are constantly evolving as tumor cells face changing selective pressures induced by the cells themselves, the microenvironment and drug treatments. Tumors are also complex ecosystems where different, sometime heterogeneous, subclonal tumor populations and a variety of nontumor cells coexist in a constantly evolving manner. The interactions between molecules and between cells that arise as a result of these alterations and ecosystems are even more complex. The cancer research community is increasingly embracing this complexity and adopting a combination of systems biology methods and integrated analyses to understand and predictively model the activity of cancer cells. Systems biology approaches are helping to understand the mechanisms of tumor progression and design more effective cancer therapies. These approaches work in tandem with rapid technological advancements that enable data acquisition on a broader scale, with finer accuracy, higher dimensionality and higher throughput than ever. Using such data, computational and mathematical models help identify key deregulated functions and processes, establish predictive biomarkers and optimize therapeutic strategies. Moving forward, implementing patient-specific computational and mathematical models of cancer will significantly improve the specificity and efficacy of targeted therapy, and will accelerate the adoption of personalized and precision cancer medicine.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Negrini S, Gorgoulis VG, Halazonetis TD . Genomic instability—an evolving hallmark of cancer. Nat Rev Mol Cell Biol 2010; 11: 220–228.
Shih AH, Abdel-Wahab O, Patel JP, Levine RL . The role of mutations in epigenetic regulators in myeloid malignancies. Nat Rev Cancer 2012; 12: 599–612.
Vogelstein B, Papadopoulos N, Velculescu VE, Zhou S, Diaz LA Jr., Kinzler KW . Cancer genome landscapes. Science 2013; 339: 1546–1558.
Suva ML, Riggi N, Bernstein BE . Epigenetic reprogramming in cancer. Science 2013; 339: 1567–1570.
Quail DF, Joyce JA . Microenvironmental regulation of tumor progression and metastasis. Nat Med 2013; 19: 1423–1437.
Greaves M, Maley CC . Clonal evolution in cancer. Nature 2012; 481: 306–313.
Hay M, Thomas DW, Craighead JL, Economides C, Rosenthal J . Clinical development success rates for investigational drugs. Nat Biotechnol 2014; 32: 40–51.
Johannessen CM, Boehm JS, Kim SY, Thomas SR, Wardwell L, Johnson LA et al. COT drives resistance to RAF inhibition through MAP kinase pathway reactivation. Nature 2010; 468: 968–972.
Poulikakos PI, Persaud Y, Janakiraman M, Kong X, Ng C, Moriceau G et al. RAF inhibitor resistance is mediated by dimerization of aberrantly spliced BRAF(V600E). Nature 2011; 480: 387–390.
Emery CM, Vijayendran KG, Zipser MC, Sawyer AM, Niu L, Kim JJ et al. MEK1 mutations confer resistance to MEK and B-RAF inhibition. Proc Natl Acad Sci USA 2009; 106: 20411–20416.
The Cancer Genome Atlas Network. Comprehensive molecular characterization of clear cell renal cell carcinoma. Nature 2013; 499: 43–49.
Finak G, Bertos N, Pepin F, Sadekova S, Souleimanova M, Zhao H et al. Stromal gene expression predicts clinical outcome in breast cancer. Nat Med 2008; 14: 518–527.
Casado P, Rodriguez-Prados JC, Cosulich SC, Guichard S, Vanhaesebroeck B, Joel S et al. Kinase-substrate enrichment analysis provides insights into the heterogeneity of signaling pathway activation in leukemia cells. Sci Signal 2013; 6: rs6.
Niepel M, Hafner M, Pace EA, Chung M, Chai DH, Zhou L et al. Profiles of basal and stimulated receptor signaling networks predict drug response in breast cancer lines. Sci Signal 2013; 6: ra84.
Kirouac DC, Du JY, Lahdenranta J, Overland R, Yarar D, Paragas V et al. Computational modeling of ERBB2-amplified breast cancer identifies combined ErbB2/3 blockade as superior to the combination of MEK and AKT inhibitors. Sci Signal 2013; 6: ra68.
Klinger B, Sieber A, Fritsche-Guenther R, Witzel F, Berry L, Schumacher D et al. Network quantification of EGFR signaling unveils potential for targeted combination therapy. Mol Syst Biol 2013; 9: 673.
Faratian D, Goltsov A, Lebedeva G, Sorokin A, Moodie S, Mullen P et al. Systems biology reveals new strategies for personalizing cancer medicine and confirms the role of PTEN in resistance to trastuzumab. Cancer Res 2009; 69: 6713–6720.
Leder K, Pitter K, Laplant Q, Hambardzumyan D, Ross BD, Chan TA et al. Mathematical modeling of PDGF-driven glioblastoma reveals optimized radiation dosing schedules. Cell 2014; 156: 603–616.
Almendro V, Cheng YK, Randles A, Itzkovitz S, Marusyk A, Ametller E et al. Inference of tumor evolution during chemotherapy by computational modeling and in situ analysis of genetic and phenotypic cellular diversity. Cell Rep 2014; 6: 514–527.
Landan G, Cohen NM, Mukamel Z, Bar A, Molchadsky A, Brosh R et al. Epigenetic polymorphism and the stochastic formation of differentially methylated regions in normal and cancerous tissues. Nat Genet 2012; 44: 1207–1214.
Choi JD, Lee JS . Interplay between epigenetics and genetics in cancer. Genomics Inform 2013; 11: 164–173.
Ehrlich M . DNA methylation in cancer: too much, but also too little. Oncogene 2002; 21: 5400–5413.
Seligson DB, Horvath S, Shi T, Yu H, Tze S, Grunstein M et al. Global histone modification patterns predict risk of prostate cancer recurrence. Nature 2005; 435: 1262–1266.
Pan H, Jiang Y, Redmond D, Nie K, Cerchietti L, Shaknovich R et al. Epigenomic evolution in diffuse large B-cell lymphomas. Blood 2013; 122: 634–634.
Tamborero D, Gonzalez-Perez A, Perez-Llamas C, Deu-Pons J, Kandoth C, Reimand J et al. Comprehensive identification of mutational cancer driver genes across 12 tumor types. Sci Rep 2013; 3: 2650.
Ashworth A, Lord CJ, Reis-Filho JS . Genetic interactions in cancer progression and treatment. Cell 2011; 145: 30–38.
Muller FL, Colla S, Aquilanti E, Manzo VE, Genovese G, Lee J et al. Passenger deletions generate therapeutic vulnerabilities in cancer. Nature 2012; 488: 337–342.
McFarland CD, Korolev KS, Kryukov GV, Sunyaev SR, Mirny LA . Impact of deleterious passenger mutations on cancer progression. Proc Natl Acad Sci USA 2013; 110: 2910–2915.
Nowell PC . The clonal evolution of tumor cell populations. Science 1976; 194: 23–28.
Muller PA, Vousden KH . p53 mutations in cancer. Nat Cell Biol 2013; 15: 2–8.
Lee W, Jiang Z, Liu J, Haverty PM, Guan Y, Stinson J et al. The mutation spectrum revealed by paired genome sequences from a lung cancer patient. Nature 2010; 465: 473–477.
Berger MF, Hodis E, Heffernan TP, Deribe YL, Lawrence MS, Protopopov A et al. Melanoma genome sequencing reveals frequent PREX2 mutations. Nature 2012; 485: 502–506.
Poon SL, Pang ST, McPherson JR, Yu W, Huang KK, Guan P et al. Genome-wide mutational signatures of aristolochic acid and its application as a screening tool. Sci Transl Med 2013; 5: 197ra101.
Marusyk A, Almendro V, Polyak K . Intra-tumour heterogeneity: a looking glass for cancer? Nat Rev Cancer 2012; 12: 323–334.
Gerlinger M, Rowan AJ, Horswell S, Larkin J, Endesfelder D, Gronroos E et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N Engl J Med 2012; 366: 883–892.
Sottoriva A, Spiteri I, Piccirillo SG, Touloumis A, Collins VP, Marioni JC et al. Intratumor heterogeneity in human glioblastoma reflects cancer evolutionary dynamics. Proc Natl Acad Sci USA 2013; 110: 4009–4014.
Bashashati A, Ha G, Tone A, Ding J, Prentice LM, Roth A et al. Distinct evolutionary trajectories of primary high-grade serous ovarian cancers revealed through spatial mutational profiling. J Pathol 2013; 231: 21–34.
Gerlinger M, Horswell S, Larkin J, Rowan AJ, Salm MP, Varela I et al. Genomic architecture and evolution of clear cell renal cell carcinomas defined by multiregion sequencing. Nat Genet 2014; 46: 225–233.
Anderson K, Lutz C, van Delft FW, Bateman CM, Guo Y, Colman SM et al. Genetic variegation of clonal architecture and propagating cells in leukaemia. Nature 2011; 469: 356–361.
Korolev KS, Xavier JB, Gore J . Turning ecology and evolution against cancer. Nat Rev Cancer 2014; 14: 371–380.
Cleary AS, Leonard TL, Gestl SA, Gunther EJ . Tumour cell heterogeneity maintained by cooperating subclones in Wnt-driven mammary cancers. Nature 2014; 508: 113–117.
Campbell PJ, Yachida S, Mudie LJ, Stephens PJ, Pleasance ED, Stebbings LA et al. The patterns and dynamics of genomic instability in metastatic pancreatic cancer. Nature 2010; 467: 1109–1113.
Lohr JG, Stojanov P, Carter SL, Cruz-Gordillo P, Lawrence MS, Auclair D et al. Widespread genetic heterogeneity in multiple myeloma: implications for targeted therapy. Cancer Cell 2014; 25: 91–101.
Landau DA, Carter SL, Stojanov P, McKenna A, Stevenson K, Lawrence MS et al. Evolution and impact of subclonal mutations in chronic lymphocytic leukemia. Cell 2013; 152: 714–726.
Johnson BE, Mazor T, Hong C, Barnes M, Aihara K, McLean CY et al. Mutational analysis reveals the origin and therapy-driven evolution of recurrent glioma. Science 2014; 343: 189–193.
Kreso A, O'Brien CA, van Galen P, Gan OI, Notta F, Brown AM et al. Variable clonal repopulation dynamics influence chemotherapy response in colorectal cancer. Science 2013; 339: 543–548.
Wargo AR, Huijben S, de Roode JC, Shepherd J, Read AF . Competitive release and facilitation of drug-resistant parasites after therapeutic chemotherapy in a rodent malaria model. Proc Natl Acad Sci USA 2007; 104: 19914–19919.
Ding L, Ley TJ, Larson DE, Miller CA, Koboldt DC, Welch JS et al. Clonal evolution in relapsed acute myeloid leukaemia revealed by whole-genome sequencing. Nature 2012; 481: 506–510.
Metzker ML . Sequencing technologies - the next generation. Nat Rev Genet 2010; 11: 31–46.
Ley TJ, Mardis ER, Ding L, Fulton B, McLellan MD, Chen K et al. DNA sequencing of a cytogenetically normal acute myeloid leukaemia genome. Nature 2008; 456: 66–72.
Ojesina AI, Lichtenstein L, Freeman SS, Pedamallu CS, Imaz-Rosshandler I, Pugh TJ et al. Landscape of genomic alterations in cervical carcinomas. Nature 2014; 506: 371–375.
Baca SC, Prandi D, Lawrence MS, Mosquera JM, Romanel A, Drier Y et al. Punctuated evolution of prostate cancer genomes. Cell 2013; 153: 666–677.
Sjoblom T, Jones S, Wood LD, Parsons DW, Lin J, Barber TD et al. The consensus coding sequences of human breast and colorectal cancers. Science 2006; 314: 268–274.
Jones S, Hruban RH, Kamiyama M, Borges M, Zhang X, Parsons DW et al. Exomic sequencing identifies PALB2 as a pancreatic cancer susceptibility gene. Science 2009; 324: 217.
Frampton GM, Fichtenholtz A, Otto GA, Wang K, Downing SR, He J et al. Development and validation of a clinical cancer genomic profiling test based on massively parallel DNA sequencing. Nat Biotechnol 2013; 31: 1023–1031.
Pike BL, Greiner TC, Wang X, Weisenburger DD, Hsu YH, Renaud G et al. DNA methylation profiles in diffuse large B-cell lymphoma and their relationship to gene expression status. Leukemia 2008; 22: 1035–1043.
Kim JH, Dhanasekaran SM, Prensner JR, Cao X, Robinson D, Kalyana-Sundaram S et al. Deep sequencing reveals distinct patterns of DNA methylation in prostate cancer. Genome Res 2011; 21: 1028–1041.
Berman BP, Weisenberger DJ, Aman JF, Hinoue T, Ramjan Z, Liu Y et al. Regions of focal DNA hypermethylation and long-range hypomethylation in colorectal cancer coincide with nuclear lamina-associated domains. Nat Genet 2012; 44: 40–46.
Hon GC, Hawkins RD, Caballero OL, Lo C, Lister R, Pelizzola M et al. Global DNA hypomethylation coupled to repressive chromatin domain formation and gene silencing in breast cancer. Genome Res 2012; 22: 246–258.
Hansen KD, Timp W, Bravo HC, Sabunciyan S, Langmead B, McDonald OG et al. Increased methylation variation in epigenetic domains across cancer types. Nat Genet 2011; 43: 768–775.
Booth MJ, Branco MR, Ficz G, Oxley D, Krueger F, Reik W et al. Quantitative sequencing of 5-methylcytosine and 5-hydroxymethylcytosine at single-base resolution. Science 2012; 336: 934–937.
Yang H, Liu Y, Bai F, Zhang JY, Ma SH, Liu J et al. Tumor development is associated with decrease of TET gene expression and 5-methylcytosine hydroxylation. Oncogene 2013; 32: 663–669.
Eswaran J, Cyanam D, Mudvari P, Reddy SD, Pakala SB, Nair SS et al. Transcriptomic landscape of breast cancers through mRNA sequencing. Sci Rep 2012; 2: 264.
Horvath A, Pakala SB, Mudvari P, Reddy SD, Ohshiro K, Casimiro S et al. Novel insights into breast cancer genetic variance through RNA sequencing. Sci Rep 2013; 3: 2256.
Maher CA, Kumar-Sinha C, Cao X, Kalyana-Sundaram S, Han B, Jing X et al. Transcriptome sequencing to detect gene fusions in cancer. Nature 2009; 458: 97–101.
Eswaran J, Horvath A, Godbole S, Reddy SD, Mudvari P, Ohshiro K et al. RNA sequencing of cancer reveals novel splicing alterations. Sci Rep 2013; 3: 1689.
Navin N, Kendall J, Troge J, Andrews P, Rodgers L, McIndoo J et al. Tumour evolution inferred by single-cell sequencing. Nature 2011; 472: 90–94.
Xu X, Hou Y, Yin X, Bao L, Tang A, Song L et al. Single-cell exome sequencing reveals single-nucleotide mutation characteristics of a kidney tumor. Cell 2012; 148: 886–895.
Hou Y, Song L, Zhu P, Zhang B, Tao Y, Xu X et al. Single-cell exome sequencing and monoclonal evolution of a JAK2-negative myeloproliferative neoplasm. Cell 2012; 148: 873–885.
Dalerba P, Kalisky T, Sahoo D, Rajendran PS, Rothenberg ME, Leyrat AA et al. Single-cell dissection of transcriptional heterogeneity in human colon tumors. Nat Biotechnol 2011; 29: 1120–1127.
Stephens PJ, Greenman CD, Fu B, Yang F, Bignell GR, Mudie LJ et al. Massive genomic rearrangement acquired in a single catastrophic event during cancer development. Cell 2011; 144: 27–40.
Vandin F, Upfal E, Raphael BJ . Algorithms for detecting significantly mutated pathways in cancer. J Comput Biol 2011; 18: 507–522.
Vaske CJ, Benz SC, Sanborn JZ, Earl D, Szeto C, Zhu J et al. Inference of patient-specific pathway activities from multi-dimensional cancer genomics data using PARADIGM. Bioinformatics 2010; 26: i237–i245.
Hu J, Locasale JW, Bielas JH, O'Sullivan J, Sheahan K, Cantley LC et al. Heterogeneity of tumor-induced gene expression changes in the human metabolic network. Nat Biotechnol 2013; 31: 522–529.
Ciriello G, Miller ML, Aksoy BA, Senbabaoglu Y, Schultz N, Sander C . Emerging landscape of oncogenic signatures across human cancers. Nat Genet 2013; 45: 1127–1133.
Alexandrov LB, Nik-Zainal S, Wedge DC, Aparicio SA, Behjati S, Biankin AV et al. Signatures of mutational processes in human cancer. Nature 2013; 500: 415–421.
Kandoth C, Schultz N, Cherniack AD, Akbani R, Liu Y, Shen H et al. Integrated genomic characterization of endometrial carcinoma. Nature 2013; 497: 67–73.
Alizadeh AA, Eisen MB, Davis RE, Ma C, Lossos IS, Rosenwald A et al. Distinct types of diffuse large B-cell lymphoma identified by gene expression profiling. Nature 2000; 403: 503–511.
Wright G, Tan B, Rosenwald A, Hurt EH, Wiestner A, Staudt LM . A gene expression-based method to diagnose clinically distinct subgroups of diffuse large B cell lymphoma. Proc Natl Acad Sci USA 2003; 100: 9991–9996.
Keutgen XM, Filicori F, Crowley MJ, Wang Y, Scognamiglio T, Hoda R et al. A panel of four miRNAs accurately differentiates malignant from benign indeterminate thyroid lesions on fine needle aspiration. Clin Cancer Res 2012; 18: 2032–2038.
Cheng WY, Ou Yang TH, Anastassiou D . Development of a prognostic model for breast cancer survival in an open challenge environment. Sci Transl Med 2013; 5: 181ra150.
Margolin AA, Bilal E, Huang E, Norman TC, Ottestad L, Mecham BH et al. Systematic analysis of challenge-driven improvements in molecular prognostic models for breast cancer. Sci Transl Med 2013; 5: 181re181.
van 't Veer LJ, Dai H, van de Vijver MJ, He YD, Hart AA, Mao M et al. Gene expression profiling predicts clinical outcome of breast cancer. Nature 2002; 415: 530–536.
Basanta D, Gatenby RA, Anderson AR . Exploiting evolution to treat drug resistance: combination therapy and the double bind. Mol Pharm 2012; 9: 914–921.
Reth M, Brummer T . Feedback regulation of lymphocyte signalling. Nat Rev Immunol 2004; 4: 269–277.
Vogelstein B, Kinzler KW . Cancer genes and the pathways they control. Nat Med 2004; 10: 789–799.
Burstein HJ . The distinctive nature of HER2-positive breast cancers. N Engl J Med 2005; 353: 1652–1654.
Davies H, Bignell GR, Cox C, Stephens P, Edkins S, Clegg S et al. Mutations of the BRAF gene in human cancer. Nature 2002; 417: 949–954.
Li J, Yen C, Liaw D, Podsypanina K, Bose S, Wang SI et al. PTEN, a putative protein tyrosine phosphatase gene mutated in human brain, breast, and prostate cancer. Science 1997; 275: 1943–1947.
Pierobon M, Wulfkuhle J, Liotta L, Petricoin E . Application of molecular technologies for phosphoproteomic analysis of clinical samples. Oncogene (e-pub ahead of print 10 March 2014; doi:10.1038/onc.2014.16).
Olsen JV, Blagoev B, Gnad F, Macek B, Kumar C, Mortensen P et al. Global, in vivo, and site-specific phosphorylation dynamics in signaling networks. Cell 2006; 127: 635–648.
Moritz A, Li Y, Guo A, Villen J, Wang Y, MacNeill J et al. Akt-RSK-S6 kinase signaling networks activated by oncogenic receptor tyrosine kinases. Sci Signal 2010; 3: ra64.
Bendall SC, Simonds EF, Qiu P, Amir el AD, Krutzik PO, Finck R et al. Single-cell mass cytometry of differential immune and drug responses across a human hematopoietic continuum. Science 2011; 332: 687–696.
Spurrier B, Ramalingam S, Nishizuka S . Reverse-phase protein lysate microarrays for cell signaling analysis. Nat Protoc 2008; 3: 1796–1808.
The Cancer Genome Atlas Network. Comprehensive molecular portraits of human breast tumours. Nature 2012; 490: 61–70.
Mao M, Tian F, Mariadason JM, Tsao CC, Lemos R Jr. Dayyani F et al. Resistance to BRAF inhibition in BRAF-mutant colon cancer can be overcome with PI3K inhibition or demethylating agents. Clin Cancer Res 2013; 19: 657–667.
Greger JG, Eastman SD, Zhang V, Bleam MR, Hughes AM, Smitheman KN et al. Combinations of BRAF, MEK, and PI3K/mTOR inhibitors overcome acquired resistance to the BRAF inhibitor GSK2118436 dabrafenib, mediated by NRAS or MEK mutations. Mol Cancer Ther 2012; 11: 909–920.
Flaherty KT, Infante JR, Daud A, Gonzalez R, Kefford RF, Sosman J et al. Combined BRAF and MEK inhibition in melanoma with BRAF V600 mutations. N Engl J Med 2012; 367: 1694–1703.
Wagner JP, Wolf-Yadlin A, Sevecka M, Grenier JK, Root DE, Lauffenburger DA et al. Receptor tyrosine kinases fall into distinct classes based on their inferred signaling networks. Sci Signal 2013; 6: ra58.
Lu Y, Muller M, Smith D, Dutta B, Komurov K, Iadevaia S et al. Kinome siRNA-phosphoproteomic screen identifies networks regulating AKT signaling. Oncogene 2011; 30: 4567–4577.
Liotta LA, Kohn EC . The microenvironment of the tumour-host interface. Nature 2001; 411: 375–379.
De Palma M, Lewis CE . Macrophage regulation of tumor responses to anticancer therapies. Cancer Cell 2013; 23: 277–286.
Pollard JW . Tumour-educated macrophages promote tumour progression and metastasis. Nat Rev Cancer 2004; 4: 71–78.
Derynck R, Akhurst RJ, Balmain A . TGF-beta signaling in tumor suppression and cancer progression. Nat Genet 2001; 29: 117–129.
Paez-Ribes M, Allen E, Hudock J, Takeda T, Okuyama H, Vinals F et al. Antiangiogenic therapy elicits malignant progression of tumors to increased local invasion and distant metastasis. Cancer Cell 2009; 15: 220–231.
Lu KV, Chang JP, Parachoniak CA, Pandika MM, Aghi MK, Meyronet D et al. VEGF inhibits tumor cell invasion and mesenchymal transition through a MET/VEGFR2 complex. Cancer Cell 2012; 22: 21–35.
Sheehan KM, Gulmann C, Eichler GS, Weinstein JN, Barrett HL, Kay EW et al. Signal pathway profiling of epithelial and stromal compartments of colonic carcinoma reveals epithelial-mesenchymal transition. Oncogene 2008; 27: 323–331.
Kahlert C, Pecqueux M, Halama N, Dienemann H, Muley T, Pfannschmidt J et al. Tumour-site-dependent expression profile of angiogenic factors in tumour-associated stroma of primary colorectal cancer and metastases. Br J Cancer 2014; 110: 441–449.
Park ES, Kim SJ, Kim SW, Yoon SL, Leem SH, Kim SB et al. Cross-species hybridization of microarrays for studying tumor transcriptome of brain metastasis. Proc Natl Acad Sci USA 2011; 108: 17456–17461.
Wu Y, Zhang W, Li J, Zhang Y . Optical imaging of tumor microenvironment. Am J Nucl Med Mol Imaging 2013; 3: 1–15.
Nakasone ES, Askautrud HA, Kees T, Park JH, Plaks V, Ewald AJ et al. Imaging tumor-stroma interactions during chemotherapy reveals contributions of the microenvironment to resistance. Cancer Cell 2012; 21: 488–503.
Wang Y, Zhou K, Huang G, Hensley C, Huang X, Ma X et al. A nanoparticle-based strategy for the imaging of a broad range of tumours by nonlinear amplification of microenvironment signals. Nat Mater 2014; 13: 204–212.
Vakoc BJ, Lanning RM, Tyrrell JA, Padera TP, Bartlett LA, Stylianopoulos T et al. Three-dimensional microscopy of the tumor microenvironment in vivo using optical frequency domain imaging. Nat Med 2009; 15: 1219–1223.
Straussman R, Morikawa T, Shee K, Barzily-Rokni M, Qian ZR, Du J et al. Tumour micro-environment elicits innate resistance to RAF inhibitors through HGF secretion. Nature 2012; 487: 500–504.
Kenny HA, Krausz T, Yamada SD, Lengyel E . Use of a novel 3D culture model to elucidate the role of mesothelial cells, fibroblasts and extra-cellular matrices on adhesion and invasion of ovarian cancer cells to the omentum. Int J Cancer 2007; 121: 1463–1472.
Wu Y, Lu Y, Chen W, Fu J, Fan R . In silico experimentation of glioma microenvironment development and anti-tumor therapy. PLoS Comput Biol 2012; 8: e1002355.
Eikenberry S, Thalhauser C, Kuang Y . Tumor-immune interaction, surgical treatment, and cancer recurrence in a mathematical model of melanoma. PLoS Comput Biol 2009; 5: e1000362.
Oved K, Eden E, Akerman M, Noy R, Wolchinsky R, Izhaki O et al. Predicting and controlling the reactivity of immune cell populations against cancer. Mol Syst Biol 2009; 5: 265.
Venkatasubramanian R, Arenas RB, Henson MA, Forbes NS . Mechanistic modelling of dynamic MRI data predicts that tumour heterogeneity decreases therapeutic response. Br J Cancer 2010; 103: 486–497.
Choe SC, Zhao G, Zhao Z, Rosenblatt JD, Cho HM, Shin SU et al. Model for in vivo progression of tumors based on co-evolving cell population and vasculature. Sci Rep 2011; 1: 31.
Harris TJ, McCormick F . The molecular pathology of cancer. Nat Rev Clin Oncol 2010; 7: 251–265.
Hanahan D, Weinberg RA . Hallmarks of cancer: the next generation. Cell 2011; 144: 646–674.
Wistuba II, Gelovani JG, Jacoby JJ, Davis SE, Herbst RS . Methodological and practical challenges for personalized cancer therapies. Nat Rev Clin Oncol 2011; 8: 135–141.
Piccart-Gebhart MJ, Procter M, Leyland-Jones B, Goldhirsch A, Untch M, Smith I et al. Trastuzumab after adjuvant chemotherapy in HER2-positive breast cancer. N Engl J Med 2005; 353: 1659–1672.
Druker BJ, Talpaz M, Resta DJ, Peng B, Buchdunger E, Ford JM et al. Efficacy and safety of a specific inhibitor of the BCR-ABL tyrosine kinase in chronic myeloid leukemia. N Engl J Med 2001; 344: 1031–1037.
Maemondo M, Inoue A, Kobayashi K, Sugawara S, Oizumi S, Isobe H et al. Gefitinib or chemotherapy for non-small-cell lung cancer with mutated EGFR. N Engl J Med 2010; 362: 2380–2388.
Chapman PB, Hauschild A, Robert C, Haanen JB, Ascierto P, Larkin J et al. Improved survival with vemurafenib in melanoma with BRAF V600E mutation. N Engl J Med 2011; 364: 2507–2516.
Druker BJ, Guilhot F, O'Brien SG, Gathmann I, Kantarjian H, Gattermann N et al. Five-year follow-up of patients receiving imatinib for chronic myeloid leukemia. N Engl J Med 2006; 355: 2408–2417.
Slamon DJ, Leyland-Jones B, Shak S, Fuchs H, Paton V, Bajamonde A et al. Use of chemotherapy plus a monoclonal antibody against HER2 for metastatic breast cancer that overexpresses HER2. N Engl J Med 2001; 344: 783–792.
Van Allen EM, Wagle N, Stojanov P, Perrin DL, Cibulskis K, Marlow S et al. Whole-exome sequencing and clinical interpretation of formalin-fixed, paraffin-embedded tumor samples to guide precision cancer medicine. Nat Med 2014; 20: 682–688.
Barretina J, Caponigro G, Stransky N, Venkatesan K, Margolin AA, Kim S et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 2012; 483: 603–607.
Lamb J, Crawford ED, Peck D, Modell JW, Blat IC, Wrobel MJ et al. The Connectivity Map: using gene-expression signatures to connect small molecules, genes, and disease. Science 2006; 313: 1929–1935.
Jahchan NS, Dudley JT, Mazur PK, Flores N, Yang D, Palmerton A et al. A drug repositioning approach identifies tricyclic antidepressants as inhibitors of small cell lung cancer and other neuroendocrine tumors. Cancer Discov 2013; 3: 1364–1377.
Chong CR, Janne PA . The quest to overcome resistance to EGFR-targeted therapies in cancer. Nat Med 2013; 19: 1389–1400.
Roychowdhury S, Iyer MK, Robinson DR, Lonigro RJ, Wu YM, Cao X et al. Personalized oncology through integrative high-throughput sequencing: a pilot study. Sci Transl Med 2011; 3: 111ra121.
Costello JC, Heiser LM, Georgii E, Gonen M, Menden MP, Wang NJ et al. A community effort to assess and improve drug sensitivity prediction algorithms. Nat Biotechnol 2014.
Bild AH, Yao G, Chang JT, Wang Q, Potti A, Chasse D et al. Oncogenic pathway signatures in human cancers as a guide to targeted therapies. Nature 2006; 439: 353–357.
Wang K, Saito M, Bisikirska BC, Alvarez MJ, Lim WK, Rajbhandari P et al. Genome-wide identification of post-translational modulators of transcription factor activity in human B cells. Nat Biotechnol 2009; 27: 829–839.
Karr JR, Sanghvi JC, Macklin DN, Gutschow MV, Jacobs JM, Bolival B Jr. et al. A whole-cell computational model predicts phenotype from genotype. Cell 2012; 150: 389–401.
Acknowledgements
We gratefully acknowledge insightful comments from Dr Yanwen Jiang (Elemento lab) and our anonymous reviewer. We also thank other members in the laboratory and Physiology, Biophysics and Systems Biology program of Weill Cornell Graduate School for helpful discussions. This work was supported by NIH, CAREER Grant from National Science Foundation, Starr Cancer Consortium and Hirschl Trust.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Du, W., Elemento, O. Cancer systems biology: embracing complexity to develop better anticancer therapeutic strategies. Oncogene 34, 3215–3225 (2015). https://doi.org/10.1038/onc.2014.291
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2014.291
This article is cited by
-
Adverse effects of tyrosine kinase inhibitors in cancer therapy: pathophysiology, mechanisms and clinical management
Signal Transduction and Targeted Therapy (2023)
-
The network structure of hematopoietic cancers
Scientific Reports (2023)
-
Bifurcation analysis of a tuberculosis progression model for drug target identification
Scientific Reports (2023)
-
Natural Products in Cancer Therapy: Past, Present and Future
Natural Products and Bioprospecting (2021)